Abstract
Background: Head and neck cancer (HNC) is one of the most prevalent and challenging diseases affecting a large population worldwide. Functional genomics can help understand the disease, but expressed gene therapy is uncertain. The study sought to identify specific genetic mutations and protein expression profiles in HNC.
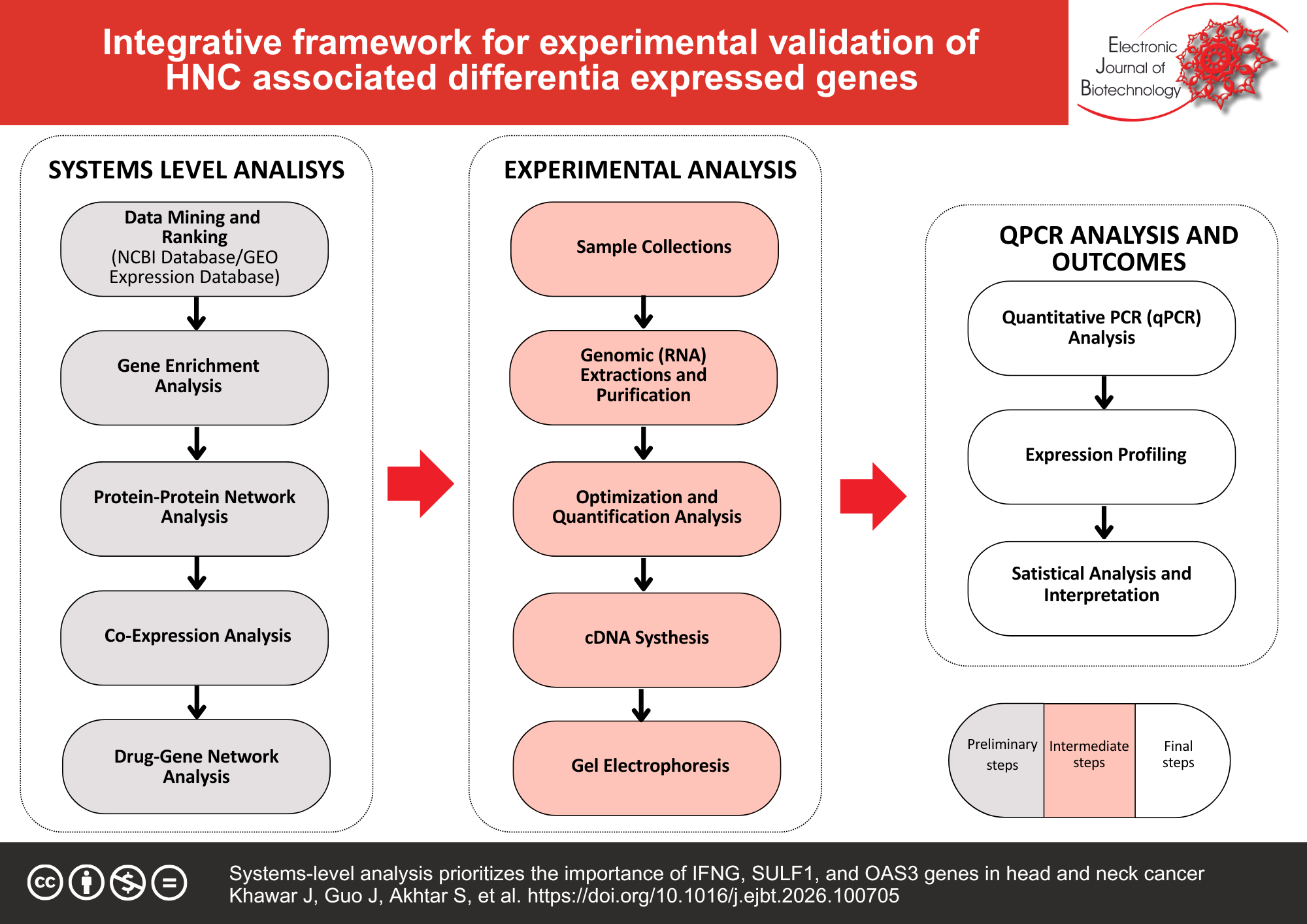
Results: We ranked IFNG, SULF1, and OAS3 as three HNC-related genes (p < 0.05) based on the data mining. N-acetyleglucosamine-6-sulfatase activity, arylsulfatase, 2,5-oligoadenylate synthetase activity, interferon-gamma receptor binding, and other essential biological processes were all significantly correlated with the gene ontology (GO) terms. Nucleotide excision repair pathways, RNA polymerase-I transcription start and termination, RNA polymerase-II promoter escape, pyrimidine biosynthesis, and interferon-gamma signaling were all linked in the pathway enrichment. OAS1, IFIT1, CD4, STAT3, NFKBIA, RIPK1, SLCO5A1, and others are functionally connected to the co-expressed genes, while COL3A1 and SCEL are indirectly linked. Compared to controls, the quantitative PCR (qPCR) of these genes showed a significant two-fold change (FC) expression (2−DDCT) pattern of SULF1 (FC ≤ 1.2), OAS3 (FC ≤ 0.13), and IFNG (FC ≤ 0.12) compared to reference gene GAPDH (FC = 1). Pathophysiological cancer development is associated with up- and downregulated expression of these genes. The study found that personalized medicine can improve HNC treatment by adapting medication to each patient’s tumor’s molecular traits.
Conclusions: A substantial correlation between the pathophysiology of HNC and the IFNG, SULF1, and OAS3 genes is found. This research could expedite the progress of drug discovery and aid in modifying HNC’s treatment approaches.
References
Addala L, Pentapati CK, Thavanati PR, et al. Risk factor profiles of head and neck cancer patients of Andhra Pradesh, India. Indian Journal of Cancer 2012;49(2):215-219. https://doi.org/10.4103/0019-509X.102865 PMid: 23107973
Mody MD, Rocco JW, Yom SS, et al. Head and neck cancer. The Lancet 2021;398(10318):2289-2299. https://doi.org/10.1016/S0140-6736(21)01550-6 PMid: 34562395
Rahman QB, Iocca O, Kufta K, et al. Global burden of head and neck cancer. Oral and Maxillofacial Surgery Clinics of North America 2020;32(3):367-375. https://doi.org/10.1016/j.coms.2020.04.002 PMid: 32482563
Datorre JG, Dos Reis MB, Sorroche BP, et al. Intratumoral Fusobacterium nucleatum is associated with better cancer-specific survival in head and neck cancer patients. Journal of Oral Microbiology 2025;17(1):2487644. https://doi.org/10.1080/20002297.2025.2487644 PMid: 40182114
Ahmad Z, Idress R, Fatima S, et al. Commonest cancers in Pakistan-findings and histopathological perspective from a premier surgical pathology center in Pakistan. Asian Pacific Journal of Cancer Prevention 2016;17(3):1061-1075. doi.org/10.7314/apjcp.2016.17.3.1061 PMid: 27039726
Dylawerska A, Barczak W, Wegner A, et al. Association of DNA repair genes polymorphisms and mutations with increased risk of head and neck cancer: a review. Medical Oncology 2017;34(12):197. https://doi.org/10.1007/s12032-017-1057-4 PMid: 29143133
Rani P, Shivaprasad, Mishra M. Emerging insights into supari (areca nut) toxicity: a review of current evidence on oral and general health. Toxin Reviews 2025;44(3):383-396. https://doi.org/10.1080/15569543.2025.2538139
Nosé V, Lazar AJ. Update from the 5th Edition of the World Health Organization Classification of Head and Neck Tumors: Familial Tumor Syndromes. Head and Neck Pathology 2022;16(1):143-157. https://doi.org/10.1007/s12105-022-01414-z PMid: 35312981
He B, Zhu R, Yang H, et al. Assessing the Impact of Data Preprocessing on Analyzing Next Generation Sequencing Data. Frontiers in Bioengineering and Biotechnology 2020;8:817. https://doi.org/10.3389/fbioe.2020.00817 PMid: 32850708
Doshi R, Hiran KK, Gök M, et al. Artificial Intelligence's Significance in Diseases with Malignant Tumours. Mesopotamian Journal of Artificial Intelligence in Healthcare 2023;2023:35-39. https://doi.org/10.58496/MJAIH/2023/007 PMid:
Huang H, Huang F, Liang X, et al. Afatinib Reverses EMT via Inhibiting CD44-Stat3 Axis to Promote Radiosensitivity in Nasopharyngeal Carcinoma. Pharmaceuticals 2022;16(1):37. https://doi.org/10.3390/ph16010037 PMid: 36678534
Pazhani J, Veeraraghavan VP, Jayaraman S. Transcription Factors: A Potential Therapeutic Target in Head and Neck Squamous Cell Carcinoma. Epigenomics 2023;15(2):57-60. https://doi.org/10.2217/epi-2023-0046 PMid:36974620
Li X, Li M, Xiang J, et al. SEPA: signaling entropy-based algorithm to evaluate personalized pathway activation for survival analysis on pan-cancer data. Bioinformatics 2022;38(9):2536-2543. https://doi.org/10.1093/bioinformatics/btac122 PMid:35199150
Garg S, Sharma N, Bharmjeet, et al. Unraveling the intricate relationship: Influence of microbiome on the host immune system in carcinogenesis. Cancer Reports 2023;6(11):e1892. https://doi.org/10.1002/cnr2.1892 PMid:37706437
Cai Y, Wu Q, Chen Y, et al. Predicting non-small cell lung cancer-related genes by a new network-based machine learning method. Frontiers in Oncology 2022;12:981154. https://doi.org/10.3389/fonc.2022.981154 PMid:36203453
Khawar J, Fatima N, Ismail M, et al. Studying association of GTF2H4, SULF1, OAS3, and IFNG genes polymorphism and risk of head and neck cancer in Southern Punjab, Pakistan. Meta Gene 2018;16:85-89. https://doi.org/10.1016/j.mgene.2018.02.002
Mzili T, Mzili M, Bouderba SI, et al. The Role of Artificial Intelligence in Early Tumor Detection: An XGBoost Risk Assessment Model for Egyptian Patients. Mesopotamian Journal of Artificial Intelligence in Healthcare 2025;2025:85-92. https://doi.org/10.58496/MJAIH/2025/008
Huang C, Liu Z, Guo Y, et al. scCancerExplorer: a comprehensive database for interactively exploring single-cell multi-omics data of human pan-cancer. Nucleic Acids Research 2025;53(D1):D1526-D1535. https://doi.org/10.1093/nar/gkae1100 PMid: 39558175
Zheng W, Li Z, Chen Y, et al. GEP-DNN4Mol: automatic chemical molecular design based on deep neural networks and gene expression programming. Health Information Science and Systems 2025;13(1):31. https://doi.org/10.1007/s13755-025-00344-8 PMid: 40144479
Asim MN, Ibrahim MA, Malik MI, et al. ADH-PPI: An attention-based deep hybrid model for protein-protein interaction prediction. iScience 2022;25(10):105169. https://doi.org/10.1016/j.isci.2022.105169 PMid: 36267921
Cui J, Yang S, Yi L, et al. Recent advances in deep learning for protein-protein interaction: a review. BioData Mining 2025;18(1):43. https://doi.org/10.1186/s13040-025-00457-6 PMid: 40524189
Singh NK, Awasthi P, Gupta A, et al. Identifying Porphyromonas gingivalis-infected hub genes and molecular mechanisms of oral squamous cell carcinoma pathogenesis. Discover Applied Sciences 2025;7:163. https://doi.org/10.1007/s42452-024-06260-y
Zheng W, Wuyun Q, Li Y, et al. Improving deep learning protein monomer and complex structure prediction using DeepMSA2 with huge metagenomics data. Nature Methods 2024;21(2):279-289. https://doi.org/10.1038/s41592-023-02130-4 PMid: 38167654
Davis AP, Wiegers TC, Johnson RJ, et al. Comparative Toxicogenomics Database (CTD): update 2023. Nucleic Acids Research 2023;51(D1):D1257-D1262. https://doi.org/10.1093/nar/gkac833 PMid: 36169237
Min D, Cheng L, Zhang F, et al. Enhancing Extracellular Electron Transfer of Shewanella oneidensis MR-1 through Coupling Improved Flavin Synthesis and Metal-Reducing Conduit for Pollutant Degradation. Environmental Science & Technology 2017;51(9):5082-5089. https://doi.org/10.1021/acs.est.6b04640 PMid: 28414427
Babicki S, Arndt D, Marcu A, et al. Heatmapper: web-enabled heat mapping for all. Nucleic Acids Research 2016;44(W1):W147-W153. https://doi.org/10.1093/nar/gkw419 PMid: 27190236
Zhang H, Jiang Y, Luo J, et al. Predictive and prognostic value of combined detection of sTim-3, PG and PD-L1 in immune checkpoint inhibitor therapy for advanced gastric cancer. American Journal of Translation Research 2024;16(11):6955-6963. https://doi.org/10.62347/MJOA5699 PMid: 39678619
Lateef RA. Machine and Deep Learning Techniques in Cancer Prediction and Risk Stratification Using Bioinformatics in Big Data Era: A Review. EDRAAK 2024;2024:118-127. https://doi.org/10.70470/EDRAAK/2024/015
Ivanenko K, Prassolov V, Khabusheva E. Transcription Factor Sp1 in the Expression of Genes Encoding Components of Mapk, JAK/STAT, and PI3K/Akt Signaling Pathways. Molecular Biology 2022;56(5):756-769. https://doi.org/10.1134/S0026893322050089 PMid: 36165020
Ko J, Humbert S, Bronson RT, et al. p35 and p39 Are Essential for Cyclin-Dependent Kinase 5 Function during Neurodevelopment. Journal of Neuroscience 2001;21(17):6758-6771. https://doi.org/10.1523/JNEUROSCI.21-17-06758.2001 PMid: 11517264
Hadia R, Singh V, Solanki N, et al. Unlocking the Clinical Significance of Cytochrome P450 Enzymes. International Journal of Pharmaceutical Investigation 2024;14(1):30-38.
Krassowski M, Paczkowska M, Cullion K, et al. ActiveDriverDB: human disease mutations and genome variation in post-translational modification sites of proteins. Nucleic Acids Research 2018;46(D1):D901-D910. https://doi.org/10.1093/nar/gkx973 PMid: 29126202
Kitts PA, Church DM, Thibaud-Nissen F, et al. Assembly: a resource for assembled genomes at NCBI. Nucleic Acids Research 2016;44(D1):D73-D80. https://doi.org/10.1093/nar/gkv1226 PMid: 26578580
Gill RM, Mehra V, Milford E, et al. Short SULF1/SULF2 splice variants predominate in mammary tumours with a potential to facilitate receptor tyrosine kinase-mediated cell signalling. Histochemistry and Cell Biology 2016;146:431-444. https://doi.org/10.1007/s00418-016-1454-3 PMid: 27294358
Brassart-Pasco S, Brézillon S, Brassart B, et al. Tumor Microenvironment: Extracellular Matrix Alterations Influence Tumor Progression. Frontiers in Oncology 2020;10:397. https://doi.org/10.3389/fonc.2020.00397 PMID 323518780
Piera-Velazquez S, Mendoza FA, Addya S, et al. Increased expression of interferon regulated and antiviral response genes in CD31+/CD102+ lung microvascular endothelial cells from systemic sclerosis patients with end-stage interstitial lung disease. Clinical and Experimental Rheumatology 2021;39(6):1298-1306. https://doi.org/10.55563/clinexprheumatol/ret1kg PMid: 33253099
Shi Y, Xu Y, Xu Z, et al. TKI resistant-based prognostic immune related gene signature in LUAD, in which FSCN1 contributes to tumor progression. Cancer Letters 2022;532:215583. https://doi.org/10.1016/j.canlet.2022.215583 PMid: 35149175
Li X, Hou L, Zhang L, et al. OAS3 is a Co-Immune Biomarker Associated With Tumour Microenvironment, Disease Staging, Prognosis, and Treatment Response in Multiple Cancer Types. Frontiers in Cell and Developmental Biology 2022;10:815480. https://doi.org/10.3389/fcell.2022.815480 PMid: 35592250
Wang Y, Hu Y, Chen L, et al. Molecular mechanisms and prognostic markers in head and neck squamous cell carcinoma: a bioinformatic analysis. International Journal of Clinical and Experimental Pathology 2020;13(3):371-381. PMid: 32269674
Kryczka J, Boncela J. Integrated Bioinformatics Analysis of the Hub Genes Involved in Irinotecan Resistance in Colorectal Cancer. Biomedicines 2022;10(7):1720. https://doi.org/10.3390/biomedicines10071720 PMid: 35885025
Ying L, Zhang F, Pan X, et al. Complement component 7 (C7), a potential tumor suppressor, is correlated with tumor progression and prognosis. Oncotarget 2016;7(52):86536-86546. https://doi.org/10.18632/oncotarget.13294 PMid: 27852032
Lu S, Peng L, Ma F, et al. Increased Expression of POSTN Predicts Poor Prognosis: a Potential Therapeutic Target for Gastric Cancer. Journal of Gastrointestinal Surgery 2023;27(2):233-249. https://doi.org/10.1007/s11605-022-05517-4 PMid: 36451060
Chae J, Choi J, Chung J. Polymeric immunoglobulin receptor (pIgR) in cancer. Journal of Cancer Research and Clinical Oncology 2023;149(19):17683-17690. https://doi.org/10.1007/s00432-023-05335-4 PMid: 37897659
Huo X, Wang S, Song H, et al. Roles of major RNA Adenosine Modifications in Head and Neck Squamous Cell Carcinoma. Frontiers in Pharmacology 2021;12:779779. https://doi.org/10.3389/fphar.2021.779779 PMid: 34899345
Ni H, Sun H, Zheng M, et al. Mining database for the expression and clinical significance of STAT family in head and neck squamous cell carcinomas. Translational Oncology 2021;14(1):100976. https://doi.org/10.1016/j.tranon.2020.100976 PMid: 33395750
Drappier M, Michiels T. Inhibition of the OAS/RNase L pathway by viruses. Current Opinion in Virology 2015;15:19-26. https://doi.org/10.1016/j.coviro.2015.07.002 PMid: 26231767
Li Y, Banerjee S, Wang Y, et al. Activation of RNase L is dependent on OAS3 expression during infection with diverse human viruses. Proceedings of the National Academy of Sciences of the United States of America 2016;113(8):2241-2246. https://doi.org/10.1073/pnas.1519657113 PMid: 26858407
Sabatini ME, Chiocca S. Human papillomavirus as a driver of head and neck cancers. British Journal of Cancer 2020;122(3):306-314. https://doi.org/10.1038/s41416-019-0602-7 PMid: 31708575
Yang E, Li MM. All About the RNA: Interferon-Stimulated Genes That Interfere with Viral RNA Processes. Frontiers in Immunology 2020;11:605024. https://doi.org/10.3389/fimmu.2020.605024 PMid: 33362792
Berinde GM, Socaciu AI, Socaciu MA, et al. Thyroid Cancer Diagnostics Related to Occupational and Environmental Risk Factors: An Integrated Risk Assessment Approach. Diagnostics 2022;12(2):318. https://doi.org/10.3390/diagnostics12020318 PMid: 35204408
Wu H, Eckhardt CM, Baccarelli AA. Molecular mechanisms of environmental exposures and human disease. Nature Reviews Genetics 2023;24(5):332-344. https://doi.org/10.1038/s41576-022-00569-3 PMid: 36717624
Meier MJ, Harrill J, Johnson K, et al. Progress in toxicogenomics to protect human health. Nature Reviews Genetics 2025;26(2):105-122. https://doi.org/10.1038/s41576-024-00767-1 PMid: 39223311
Li N, Wang J, Zhang N, et al. Cross-talk between TNF-? and IFN-? signaling in induction of B7-H1 expression in hepatocellular carcinoma cells. Cancer Immunology, Immunotherapy 2018;67(2):271-283. https://doi.org/10.1007/s00262-017-2086-8 PMid: 29090321
Zheng Y, Li Y, Lian J, et al. TNF-?-induced Tim-3 expression marks the dysfunction of infiltrating natural killer cells in human esophageal cancer. Journal of Translational Medicine 2019;17(1):165. https://doi.org/10.1186/s12967-019-1917-0 PMid: 31109341
Brasil da Costa FH, Lewis MS, Truong A, et al. SULF1 suppresses Wnt3A-driven growth of bone metastatic prostate cancer in perlecan-modified 3D cancer-stroma-macrophage triculture models. PLoS One 2020;15(5):e0230354. https://doi.org/10.1371/journal.pone.0230354 PMid: 32413029
Ouyang Q, Liu Y, Tan J, et al. Loss of ZNF587B and SULF1 contributed to cisplatin resistance in ovarian cancer cell lines based on Genome-scale CRISPR/Cas9 screening. American Journal of Cancer Research 2019;9(5):988-998. PMid: 31218106
Yang Y, Ahn J, Edwards NJ, et al. Extracellular Heparan 6-O-Endosulfatases SULF1 and SULF2 in Head and Neck Squamous Cell Carcinoma and Other Malignancies. Cancers 2022;14(22):5553. https://doi.org/10.3390/cancers14225553 PMid: 36428645
Musumeci G, Castrogiovanni P, Barbagallo I, et al. Expression of the OAS Gene Family is Highly Modulated in Subjects Affected by Juvenile Dermatomyositis, Resembling an Immune Response to a dsRNA Virus Infection. International Journal of Molecular Sciences 2018;19(9):2786. https://doi.org/10.3390/ijms19092786 PMid: 30227596
Choi UY, Kang J-S, Hwang YS, et al. Oligoadenylate synthase-like (OASL) proteins: dual functions and associations with diseases. Experimental & Molecular Medicine 2015;47(3):e144. https://doi.org/10.1038/emm.2014.110 PMid: 25744296
Shaath H, Vishnubalaji R, Elkord E, et al. Single-Cell Transcriptome Analysis Highlights a Role for Neutrophils and Inflammatory Macrophages in the Pathogenesis of Severe COVID-19. Cells 2020;9(11):2374. https://doi.org/10.3390/cells9112374 PMid: 33138195
Zhang Y, Yu C. Prognostic characterization of OAS1/OAS2/OAS3/OASL in breast cancer. BMC Cancer 2020;20(1):575. https://doi.org/10.1186/s12885-020-07034-6 PMid: 32560641

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Copyright (c) 2026 Electronic Journal of Biotechnology